
- Patent
- Our Patent solutions
- IP Intelligence SoftwareStreamline the journey from innovation to IP with our IP intelligence software, patent search, data analytics, and related services and tools.→ Read more
 Our solutionsPowerful patent searching and analysisArtificial intelligence for patent classificationGrant statistics and detailed legal status analysisSmall molecules searching and analysisDNA and amino acid searching and analysisExplore new markets and opportunitiesEasy patent searching and collaborationAnalysis from any scientific sourcesUnlock AI-powered use cases across a range of Questel software.SEP DatabaseNEWStandard Essential Patent Data & AnalyticsLatest resources
Our solutionsPowerful patent searching and analysisArtificial intelligence for patent classificationGrant statistics and detailed legal status analysisSmall molecules searching and analysisDNA and amino acid searching and analysisExplore new markets and opportunitiesEasy patent searching and collaborationAnalysis from any scientific sourcesUnlock AI-powered use cases across a range of Questel software.SEP DatabaseNEWStandard Essential Patent Data & AnalyticsLatest resources- → Questel Announces BLUESHIFT IP Has Selected Orbit Intelligence and QthenaBy adopting Questel’s solutions, the firm aims to further strengthen its ability to ...
- → AVERA-IP Selects Questel’s Qthena to Enhance AI-Driven Patent Drafting and Prosecution WorkflowsAVERA-IP Selects Questel’s Qthena to Enhance AI-Driven Patent Drafting and Prosecuti...
- → Types of Intellectual Property: Definitions, Examples, and Strategic Value Understand the key types of intellectual property—patents, trademarks, copyrights, a...
- → Questel Announces BLUESHIFT IP Has Selected Orbit Intelligence and Qthena
- IP Management SoftwareKeep your assets up to date and aligned with your corporate strategy with our Equinox suite of IP management software.→ Read more
 Our solutionsPowerful all-in-one IPMS for corporatesIPMS tailored for large corporates, built on SalesforcePowerful all-in-one IPMS for law firmsIPMS tailored for large Law FirmsCollaborative invention-to-filing workflowLatest resources
Our solutionsPowerful all-in-one IPMS for corporatesIPMS tailored for large corporates, built on SalesforcePowerful all-in-one IPMS for law firmsIPMS tailored for large Law FirmsCollaborative invention-to-filing workflowLatest resources- → How to Leverage GenAI for Advanced Patent and Trademark WorkflowsFrom invention disclosure and patent drafting to office action response management a...
- → 7 Data Security Features You Need in Your IP Management SoftwareIP management software is evolving rapidly, with vendors and users embracing the pot...
- → Does Your IP Management Technology Connect Your People, Processes, and IP Assets? To drive real efficiency across their IP management practices, portfolio owners must...
- → How to Leverage GenAI for Advanced Patent and Trademark Workflows
- AI Assistants for Patent ProductivityOptimize your entire patent process with our advanced AI assistants for patent productivity, from patent application drafting to prosecution workflow automation, claims mapping, and office action responses.→ Read more
 Our solutionsQthenaNEWAll-in-one patent prosecution workflow, data, and collaboration workspace.Patent Mapping and Claim Analysis with AIPatent Office Action Response Management with AILeverage the power of data to guide your patent prosecution with Prosecution PackIndustry-leading patent drafting software powered by generative AI.Latest resources
Our solutionsQthenaNEWAll-in-one patent prosecution workflow, data, and collaboration workspace.Patent Mapping and Claim Analysis with AIPatent Office Action Response Management with AILeverage the power of data to guide your patent prosecution with Prosecution PackIndustry-leading patent drafting software powered by generative AI.Latest resources- → Questel Announces BLUESHIFT IP Has Selected Orbit Intelligence and QthenaBy adopting Questel’s solutions, the firm aims to further strengthen its ability to ...
- → How Can AI Help IP Practitioners Manage Responses to Office Actions? The latest AI tools for IP law firms and corporations are helping IP practitioners s...
- → Generative AI and IP Practice: The New StandardPatent and trademark attorneys once viewed generative AI with skepticism. Confidenti...
- → Questel Announces BLUESHIFT IP Has Selected Orbit Intelligence and Qthena
- Patent ServicesBenefit from end-to-end support for vital but time-consuming administrative tasks with our complete range of patent support services, including searches, translations, international filings, validations, unitary patent management, recordals, renewals, and more→ Read more
 Our solutionsPatentability, Freedom-To-Operate, InvalidityPCT & national phase, Paris Convention, Unitary PatentUnitary Patent and Unified Patent Court servicesSimplified European validation servicesCentralized patent translation servicesFee and renewals managementAssignment and recordal of all IP RightsLatest resources
Our solutionsPatentability, Freedom-To-Operate, InvalidityPCT & national phase, Paris Convention, Unitary PatentUnitary Patent and Unified Patent Court servicesSimplified European validation servicesCentralized patent translation servicesFee and renewals managementAssignment and recordal of all IP RightsLatest resources- → MicroPort NeuroScientific Selects Questel for Patent Filing, Translation, Unitary Patent, and EP Validation Services By partnering with Questel, MicroPort NeuroScientific gains access to integrated IP ...
- → AI, Efficiency, and Trust: What’s Really Changing in Patent Translation? AI is reshaping patent translation, but not by replacing human expertise. It is chan...
- → PCT FILING GUIDENational phase filing requirements across 50 of the most frequently selected PCT jur...
- → MicroPort NeuroScientific Selects Questel for Patent Filing, Translation, Unitary Patent, and EP Validation Services
- Patent Strategy & Administration→ Read more
Control costs, streamline invoices, and safeguard your patent rights efficiently with our patent strategy & administration services and tools
 Our solutionsEnd-to-end IP admin, data verification and docketingInvoice review, bundling and paymentFee audits, negotiation, compliance, benchmarks and forecastingLandscaping, benchmarking, valuation and licensingDefensive PublicationBUY ONLINEPublish an invention into the public domainLatest resources
Our solutionsEnd-to-end IP admin, data verification and docketingInvoice review, bundling and paymentFee audits, negotiation, compliance, benchmarks and forecastingLandscaping, benchmarking, valuation and licensingDefensive PublicationBUY ONLINEPublish an invention into the public domainLatest resources- → Cut Intellectual Project Cost and Reduce Inefficiency: What Can In-House Legal Teams Do? There is always pressure on intellectual property costs. Indeed, an in-house legal t...
- → Centralized Patent Management Services Deliver 20% Cost Savings for Autoliv Henning Koch, IP & Patent Director at Autoliv, shares his insights and positive expe...
- → The 4 Elements of a Cost-Effective Legal DepartmentHow can a legal department reduce operational costs without sacrificing quality? Wha...
- → Cut Intellectual Project Cost and Reduce Inefficiency: What Can In-House Legal Teams Do?
- Trademark
- Our Trademark solutions
- Clearance & Watch PlatformScreen and monitor your brand and trademark rights effectively with our trademark clearance search and trademark monitoring platform→ Read more
 Our solutionsScreen trademarks with AI-driven search technologyMarkify Comprehensive SearchBUY ONLINEAI-driven trademark availability softwareBest-in-class trademark results paired with key pharmaceutical sourcesWatch for new, identical and confusingly similar trademarksMonitor marketplaces, social media, and web contentLatest resources
Our solutionsScreen trademarks with AI-driven search technologyMarkify Comprehensive SearchBUY ONLINEAI-driven trademark availability softwareBest-in-class trademark results paired with key pharmaceutical sourcesWatch for new, identical and confusingly similar trademarksMonitor marketplaces, social media, and web contentLatest resources- → TALKUAL Selects Markify Watch to Strengthen Its Brand Protection Strategy As TALKUAL continues its international growth journey, protecting and strengthening ...
- → Pointon Partners Selects Questel’s Markify platform to Screen and Monitor his Brand and Trademark Rights EffectivelyWe are excited to announce that Pointon Partners has selected Markify, our trademark...
- → How ZEISS Strengthens its Online Brand Protection and Trademark Searches with QuestelHolger Gensmantel, Head of Trademarks, shares how Questel supports the brand defense...
- → TALKUAL Selects Markify Watch to Strengthen Its Brand Protection Strategy
- IP Management SoftwareManage and maintain your trademark, design and domain name rights effectively with our state-of-the-art trademark management software→ Read more
 Our solutionsPowerful all-in-one IPMS for corporatesIPMS tailored for large corporates, built on SalesforcePowerful all-in-one IPMS for law firmsIPMS tailored for large law firmsCollaborative workflow for trademark and design proposalsLatest resources
Our solutionsPowerful all-in-one IPMS for corporatesIPMS tailored for large corporates, built on SalesforcePowerful all-in-one IPMS for law firmsIPMS tailored for large law firmsCollaborative workflow for trademark and design proposalsLatest resources- → How to Leverage GenAI for Advanced Patent and Trademark WorkflowsFrom invention disclosure and patent drafting to office action response management a...
- → 7 Data Security Features You Need in Your IP Management SoftwareIP management software is evolving rapidly, with vendors and users embracing the pot...
- → Proof of Use in Practice: Managing PoU with AI in UZ.IPProof of use often creates friction in trademark workflows. Time pressure, difficult...
- → How to Leverage GenAI for Advanced Patent and Trademark Workflows
- AI Assistants for Trademark ProductivityReview trademark search and watch results, create first drafts of goods and services descriptions, classify evidence of use, and manage office action responses effectively with Qthena, our AI assistant for trademark productivity.→ Read more
 Our solutionsQthenaNewAll-in-one digital trademark workflow automationChat with trademark search or watch results to review findings and assess riskSave time and build consistency and compliance with our AI solutionAnalyze content and draft answers with AIManage trademark evidence of use with AILatest resources
Our solutionsQthenaNewAll-in-one digital trademark workflow automationChat with trademark search or watch results to review findings and assess riskSave time and build consistency and compliance with our AI solutionAnalyze content and draft answers with AIManage trademark evidence of use with AILatest resources- → How Can AI Help IP Practitioners Manage Responses to Office Actions? The latest AI tools for IP law firms and corporations are helping IP practitioners s...
- → Murgitroyd Case Study: Significant Time Saved with Qthena and Reinvested into IP StrategiesA leading IP law firm, Murgitroyd set out to remove workflow bottlenecks, strengthen...
- → How to Leverage GenAI for Advanced Patent and Trademark WorkflowsFrom invention disclosure and patent drafting to office action response management a...
- → How Can AI Help IP Practitioners Manage Responses to Office Actions?
- Trademark, Design & Domain Services→ Read more
Access expert support throughout the complete brand lifecycle with our integrated domain name, design, and trademark solutions
 Our solutionsAnalyst-driven searches with on-demand and customizable reportingIP asset monitoring with intelligent pre-selectionCase management and take-down servicesInternational registration process supportTrademark and design renewal servicesEffective management of corporate domain namesAssignment and recordal of all IP rightsEnd-to-end IP admin, data verification, docketingLatest resources
Our solutionsAnalyst-driven searches with on-demand and customizable reportingIP asset monitoring with intelligent pre-selectionCase management and take-down servicesInternational registration process supportTrademark and design renewal servicesEffective management of corporate domain namesAssignment and recordal of all IP rightsEnd-to-end IP admin, data verification, docketingLatest resources- → Stop Searching for Registered Trademarks Online: You’re Wasting Time and Money If you only search for registered trademark rights online, you will miss many import...
- → No IP Docketing System? Keep a Clean and Efficient Legal Docket with Questel Making sure your IP legal docket is an effective resource can be a challenge, but th...
- → Managing Renewals with a Dedicated Intellectual Property Platform: How Does It Work? Traditional methods of managing IP renewals are labor-intensive and relatively high-...
- → Stop Searching for Registered Trademarks Online: You’re Wasting Time and Money
- Innovation
- Our Innovation solutions
- Innovation Management SoftwareMake your innovation management processes faster, efficient and scalable with our end-to-end innovation management platform.→ Read more
 Our solutionsBusiness-oriented dashboards for innovatorsStartup deal flow and program managementInnovation Project ManagementUnlock AI-powered use cases across a range of Questel software.Your central cockpit for your strategic AI projectsLatest resources
Our solutionsBusiness-oriented dashboards for innovatorsStartup deal flow and program managementInnovation Project ManagementUnlock AI-powered use cases across a range of Questel software.Your central cockpit for your strategic AI projectsLatest resources- → The Critical Role of Human Input in AI-Powered DecisionsWhy AI needs human input to deliver business value, and how innovation leaders are i...
- → How to Choose the Best Open Innovation Platform for Your Business NeedsYour innovation platform should provide a dynamic space where you can share ideas, t...
- → Why You Should Focus on Innovation Even at Times of Crisis Investing in innovation and innovation management is essential not only during stron...
- → The Critical Role of Human Input in AI-Powered Decisions
- Innovation ServicesSuccessful innovation processes and culture are about more than the right software tools. Our innovation services leverage proven methods and approaches to generate insights, ideas, and solutions.→ Read more
 Our solutionsMarket screening, Competitive intelligence and Technology scoutingConnect with partners, improve or localize your innovation processProducts and services namingSeamless user experience to a global audienceEngage with new audience globallyLatest resources
Our solutionsMarket screening, Competitive intelligence and Technology scoutingConnect with partners, improve or localize your innovation processProducts and services namingSeamless user experience to a global audienceEngage with new audience globallyLatest resources- → 7 Key Questions to Measure Innovation Management Success Innovation is the key to staying competitive in today’s fast-paced business world. H...
- → Software Patent: How Can It Be Done?Even though life without computers is today nearly impossible to imagine, patent-bas...
- → What is Innovation Scouting?Change is a constant. New technologies emerge regularly, reshaping product landscape...
- → 7 Key Questions to Measure Innovation Management Success
- Solutions
- Solutions
- Artificial Intelligence in IP→ Read more
Discover Questel’s holistic and responsible approach to artificial intelligence
and intellectual property and our vision to inspire the future of IP with AI Our solutionsDiscover how AI is transforming IPIndustry-leading patent drafting software powered by generative AIAI copilots for optimized patent preparation and prosecution workflowsAI copilot for automated patent classificationAI copilot for elevated trademark clearance searchUnlock AI-powered use cases across a range of Questel software.Proof of UseNewCapture Trademark Evidence of Use with AILatest resources
Our solutionsDiscover how AI is transforming IPIndustry-leading patent drafting software powered by generative AIAI copilots for optimized patent preparation and prosecution workflowsAI copilot for automated patent classificationAI copilot for elevated trademark clearance searchUnlock AI-powered use cases across a range of Questel software.Proof of UseNewCapture Trademark Evidence of Use with AILatest resources- → From Risk to Reward: What Our 2026 Industry Outlook Research Reveals About AI in IPThe latest edition of our IP Industry Outlook Research shows that AI is providing a ...
- → Generative AI and IP Practice: The New StandardPatent and trademark attorneys once viewed generative AI with skepticism. Confidenti...
- → Murgitroyd Case Study: Significant Time Saved with Qthena and Reinvested into IP StrategiesA leading IP law firm, Murgitroyd set out to remove workflow bottlenecks, strengthen...
- → From Risk to Reward: What Our 2026 Industry Outlook Research Reveals About AI in IP
- Integrated IP Ecosystem→ Read more
Optimize your intellectual property assets with our comprehensive IP portfolio management solutions,
ensuring maximum protection and strategic advantage for your business. Our solutionsIP management systems, docketing, forecasting, data analytics, blockchain, and eBilling toolsOur web-based platform for IP services managementPAVIS Connect, our solution for companies and laws firms that already have an IP management systemAccess trademark watches directly from Equinox with Markify Watch integrationLatest resources
Our solutionsIP management systems, docketing, forecasting, data analytics, blockchain, and eBilling toolsOur web-based platform for IP services managementPAVIS Connect, our solution for companies and laws firms that already have an IP management systemAccess trademark watches directly from Equinox with Markify Watch integrationLatest resources- → Types of Intellectual Property: Definitions, Examples, and Strategic Value Understand the key types of intellectual property—patents, trademarks, copyrights, a...
- → IP Valuation Services for Smarter Business DecisionsIn today’s knowledge-driven economy, intellectual property (IP) assets often hold mo...
- → Switching to a New IPMS: What Does the Data Migration Process Involve?When you’re considering switching to a new IP management system (IPMS), you’ll likel...
- → Types of Intellectual Property: Definitions, Examples, and Strategic Value
- Solutions for Law FirmsStreamline internal processes, satisfy client demand, and remain competitive with our specialist solutions for law firms→ Read moreOur solutionsAll-in-one patent prosecution workflow, data, and collaboration workspace.Patent Office Action Response Management with AIIndustry-leading patent drafting software powered by generative AIPowerful all-in-one IPMS for law firmsPowerful patent searching and analysisScreen and monitor your trademark rights effectively with our Markify platformStreamline multilingual matters with fast and reliable legal translation servicesCentralized patent translation servicesFee and renewals managementTrademark and design renewal servicesLatest resources
- → How Equinox Law Firm Supports Redchip Lawyers to Save Time and Build RelationshipsFor law firms with large, multi-disciplinary practices, an IP management system (IPM...
- → Can Your IPMS Help Build Strong Client Relationships?Cultivating strong client relationships is incredibly important for IP law firms. Yo...
- → Leverage Artificial Intelligence to Simplify Your Prosecution WorkUnlock exclusive insights and discover how Qthena, the AI-powered prosecution assist...
- → How Equinox Law Firm Supports Redchip Lawyers to Save Time and Build Relationships
- Solutions for Life Sciences→ Read more
Intellectual property management tools, services, and insights to support life sciences innovation strategies
Our solutionsDNA and amino acid searching and analysisSmall molecules searching and analysisExplore new markets and opportunitiesBest-in-class trademark results paired with key pharmaceutical sourcesPowerful all-in-one IPMS for corporatesTranslation solutions across the product lifecycle—from patent to post-marketCentralized patent translation servicesFee and renewals managementTrademark and design renewal servicesLSPN North America Awards recognize QuestelLatest resources- → Patent Data Analytics, It’s Not Just for Patent SearchingThere are many reasons why organizations may conduct patent searching exercises and—...
- → Harnessing the Power of Patent AnalyticsPatent analytics can reveal new application areas, diversification opportunities, an...
- → The Technology Landscape of Precision Medicine: An IP AnalysisTechnological developments in gene sequencing, imagery, and data analysis are enabli...
- → Patent Data Analytics, It’s Not Just for Patent Searching
- Solutions for R&D and Innovation→ Read more
Strengthen, structure, and maximize your R&D and innovation strategy, workflows, and internal processes by harnessing the latest tools and technologies, including through smart application of generative AI.
Our solutionsCreate more efficient and scalable innovation management processesCollaborative invention-to-filing workflowCollaborative worklfow for trademark and design proposalsMarket screening, competitive intelligence, and technology scoutingStreamline your research and development workflows with AILatest resources- → 3 Cutting-Edge Solutions to Support Your R&D and Innovation Strategies Discover how the latest cutting-edge software and AI-elevated processes for research...
- → 7 Key Questions to Measure Innovation Management Success Innovation is the key to staying competitive in today’s fast-paced business world. H...
- → AI Patents: What Patent Mapping Can Tell Us About Future AI Technologies Artificial intelligence (AI) is evolving all the time thanks to the growing quantiti...
- → 3 Cutting-Edge Solutions to Support Your R&D and Innovation Strategies
- Language Solutions→ Read more
Translation, interpretation and tech-enabled localization services and language solutions that help global organizations open up new markets, overcome regulatory hurdles and connect with audiences worldwide.
 Our solutionsStreamline multilingual matters with fast and reliable legal translation servicesTranslation solutions across the product lifecycle—from patent to post-marketManage operations, ensure compliance, and maintain consistent messaging worldwideLatest resources
Our solutionsStreamline multilingual matters with fast and reliable legal translation servicesTranslation solutions across the product lifecycle—from patent to post-marketManage operations, ensure compliance, and maintain consistent messaging worldwideLatest resources- → Creating Organization-Wide IP Awareness: How Leading Companies Strengthen Innovation and Reduce RiskHaving an organization wide IP Awareness is essential! But how companies can build a...
- → Employing A "Winning Strategy" For Your International MattersYear after year, in-house legal teams and outside counsel handling international and...
- → The Role Clinical Trial Translation Plays in Patient Recruitment and RetentionHealth equity is an essential goal in modern healthcare, and one pivotal aspect of a...
- → Creating Organization-Wide IP Awareness: How Leading Companies Strengthen Innovation and Reduce Risk
- Contact
- Learn & Support
- Learn and support
- Webinars & EventsAre you interested in attending one of our online or onsite event?
- Product TrainingsCustomer success is our priority. Increase your skills in the use of Questel’s software
- Product NewsA platform dedicated to software and platforms news and evolutions
- Best-in-class Customer ExperienceOur goal is to exceed our clients' expectations and share best practices
- IP TrainingIncrease the IP-IQ of your entire organization with engaging IP training programs
- Newsletter subscriptionSign up for our quarterly patent and trademark newsletters and set your email preferences below.
- Webinars & Events
- Resource HubStay up-to-date with industry best practices with our latest blogs
- Resource Hub
- About Questel
- Learn & Support
- Learn and support
- Webinars & EventsAre you interested in attending one of our online or onsite event?
- Product TrainingsCustomer success is our priority. Increase your skills in the use of Questel’s software
- Product NewsA platform dedicated to software and platforms news and evolutions
- Best-in-class Customer ExperienceOur goal is to exceed our clients' expectations and share best practices
- IP TrainingIncrease the IP-IQ of your entire organization with engaging IP training programs
- Newsletter subscriptionSign up for our quarterly patent and trademark newsletters and set your email preferences below.
- Webinars & Events
- Resource HubStay up-to-date with industry best practices with our latest blogs
- Resource Hub
- About Questel

Patent sequence search - Orbit BioSequence
OBS: From simple patent sequence search to variant analysis
DNA, RNA sequences as well as proteins have been disclosed in patents since the 60s and a few even before that. Many laws have been created and modified over time to allow different types of biological material to be patented, such as naturally occurring sequences, modified sequences, sequences used in diagnostics, sequences from plants and many other types. We have recently seen that vaccines are a hot topic and some, such as RNA vaccines, do include sequences. Industrial domains that publish sequences could seem surprising, but the food industry or detergent manufacturers for instance are some of them. Obviously, pharmaceutical industry, biotech, agrochemical and seed companies produce the bulk of sequence patents. So, why is patent sequence searching important and why is it different to other types of patent searching?
Patent Sequence Data
Starting in the 90s, and the human genome project, genomic and mRNA sequences started to become more common in patents. In some cases, whole genomes (from bacteria, fungus) which can be made up of millions of base pairs were published. Private companies disclosed and, in some cases claimed millions of short sequences. All this is happening when all patents were purely filed on paper. Electronic filing of patents and supplemental materials such as sequence listings finally became available in part due to sequence patent . Since then, we have seen an increase in the number of patents with sequences, and despite the massive rise in Chinese patents, the worldwide newly published number of patents with sequences still follows a linear curve.
 Historical trend of newly published sequence patents from 2000 to 2020 available in Orbit BioSequence.
Historical trend of newly published sequence patents from 2000 to 2020 available in Orbit BioSequence.
The historical big three authorities (USPTO, EPO and WIPO) publish their sequences. Some other authorities are very compliant such as JPO, KIPO and CIPO. Others are less systematic or stuck in the past, unfortunately. But even for highly compliant authorities, rules and laws on what sequences should be disclosed vary. It is, thus, highly recommended to have a family view of your patents since a sequence patent might be different in an EPO document than in the USPTO or WIPO documents of the same family.
Why patent sequence search is different?
Traditional IP searching is done with keywords. Since searching with keywords is imperfect, they are often combined with patent classes, synonymous lists, and many other features that, basically, attempt to alleviate the pain induced by the lack of accuracy of keywords.
Biological sequence searches are different for several reasons. First, there is a common language to describe DNA/RNA and amino acid sequences, entirely independent from the native language the patent is written in. So, no need for natural language translation.
Second, since sequences can be very long, several publication standards have existed over time to treat them separately in a sequence listing. Thus, a large majority of published sequences are simple to treat electronically. This can be contrasted with chemistry where images are still an acceptable form of publication.
Third, unless your sequence is very short, you will always want to find sequences similar to yours, not just identical. This is particularly important since small errors (OCR mistakes, publisher errors) can be somehow controlled. By contrast, if you searched for the keywords “bread yeast”, you would not find “bead yeast” even if the latter could be a spelling mistake.
Fourth, for the last 20 years or so, sequences published in a patent are numbered and referred to by the keywords SEQ ID NO. It is easy in most cases to know if, say, hit sequence 5 is claimed since it is referred to by its number in the claims section as SEQ ID NO. 5. This is a unique feature of sequences and one that is critically important, allowing us to highlight sequence instances as (claimed) for instance.
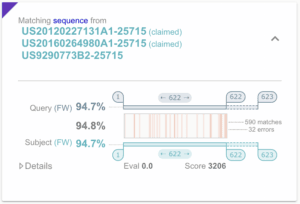
A sequence aligned to a sequence appearing in three USPTO documents, claimed or not in Orbit BioSequence.
Alignments and algorithms
Patent sequence searching consists in aligning your query sequence to sequences in a database using specific algorithms and parameters. This is all pretty complicated, however, it can be limited to a few use cases. At the risk of oversimplifying the problem, either you use a long gene sequence to find similar sequences, or you use a short sequence. For the former, everything works well, just make sure to compare your gene to both nucleotide and protein databases since you don’t know beforehand what a patent could claim or disclose. In the latter case of a short sequence, things are bit more difficult. You might want to find sequences that are perfectly matching your query or permit a few mismatches. Do you want to allow gaps? If you use 3 or 6 antibody CDRs, do you want all aligned to other CDRs, or heavy or light chains? All those questions might lead to different algorithms and parameters.But don’t worry, we have extensive documentation and a great helpdesk. Complex problems don’t always lead to simple solutions!
Pairwise vs. Variant multiple sequence alignment
Eventually, you will see pairwise alignments, in other words, your query sequence aligned to a patent sequence. This will give you intricate details of the differences between the two sequences and, combined with the available patent information, will help you decide if this alignment is relevant to your FTO, patentability, … Indeed, there can be a lot of sequences in the same patent family and many families. You will need to browse through many, but we can help with filters that will lead to only the most relevant alignments and families. However, you will miss a global view of the alignments. How many patent sequences have a lysine at position 34 of your query? This can only be done with a variant analysis.
A variant analysis will stack all patent sequences aligned to your query and will give you a global view for each query position. In other words, it will create a multiple alignment based on your query sequence. You can query, modify, and export the dataset, and most importantly, explore the variations to give you new insight into what your competitors are doing or what area are never modified for instance.

Variations at several positions using Orbit BioSequence Variant Analysis
Orbit BioSequence (OBS)
With an extensive access to patent sequences as well as non-patent sequences, Orbit BioSequence is the perfect tool for your FTO, patentability and business intelligence searches. By easily combining patent data and sequences, OBS will make your patent sequence searches a lot easier than other tools purely dedicated to sequences. Antibody and CDR, genes, primers can all be used, combined, and explored.
Interested to find out more? Contact us for specific advice or support, or watch the recording of our recent webinar Smart & visual sequence variations explorer in patent data By Orbit BioSequence.


